Institut für Visual Computing (IVC)

Gliederung
Fachbereich Informatik, Institut für Technik, Ressourcenschonung und Energieeffizienz (TREE)
Forschungsfelder
- Computational Chemistry
Standort
Sankt Augustin
Raum
A022.1
Adresse
Grantham-Allee 20
53754, Sankt Augustin
Telefon
+49 2241 865 267Profil
Die vollständigen Informationen finden Sie in der englischen Version.
Lebenslauf
Die vollständigen Informationen finden Sie in der englischen Version.
Forschungsprojekte
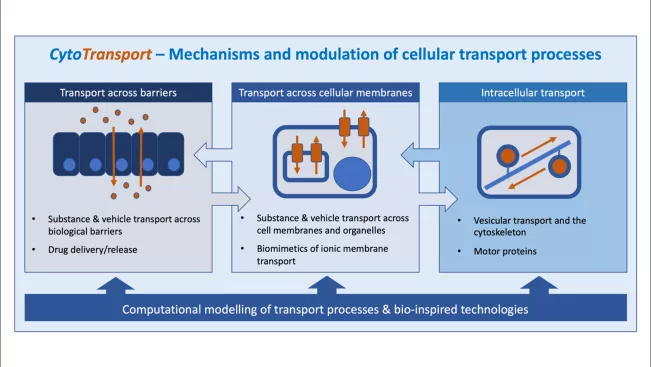
Im DFG-geförderten Forschungsverbund "CytoTransport" werden Expertisen in Biomedizin, computergestützter Modellierung, Strukturbiologie, Chemie und Materialwissenschaften vereint, um zelluläre Transportmechanismen in Gesundheit und Krankheit zu erforschen. Hierzu arbeiten 9 Arbeitsgruppen aus zwei Hochschulforschungsschwerpunkten zusammen, vertreten durch die beiden Forschungsinstitute IFGA und TREE. Ein übergeordnetes Ziel ist dabei, überlappende Interessen und Expertisen der Institute langfristig zu bündeln.
Projektleitung an der H-BRS
Prof. Dr. Mike Althaus Dr. Karl Kirschner Prof. Dr. Matthias Preller Prof. Dr. Dirk Reith Prof. Dr. Jörn Oliver Sass Prof. Dr. Margit Schulze Prof. Dr. Christopher Volk Prof. Dr. Steffen Witzleben Dr. Katrin RichterDie bio-chemische Forschung ist zunehmend auf akkurate Computermodellierung und -analyse angewiesen. Dieses Forschungsfeld ist naturgemäß hoch interdisziplinär, da physikalische Grundgesetze algorithmisch umgesetzt werden müssen, um in Anwendungen der Lebenswissenschaften relevante Beiträge liefern zu können. Das Projekt und die damit verbundene Initiative UMMBAS bündelt disziplinübergreifend die starke Expertise an der H-BRS in der Methodenentwicklung, der Visualisierung und der Anwendung computergestützter Verfahren zur Entschlüsselung materialwissenschaftlicher und biochemischer Fragestellungen.
Projektleitung an der H-BRS
Prof. Dr. Matthias PrellerIm DFG-geförderten Forschungsverbund "CytoTransport" werden Expertisen in Biomedizin, computergestützter Modellierung, Strukturbiologie, Chemie und Materialwissenschaften vereint, um zelluläre Transportmechanismen in Gesundheit und Krankheit zu erforschen. Hierzu arbeiten 9 Arbeitsgruppen aus zwei Hochschulforschungsschwerpunkten zusammen, vertreten durch die beiden Forschungsinstitute IFGA und TREE. Ein übergeordnetes Ziel ist dabei, überlappende Interessen und Expertisen der Institute langfristig zu bündeln.
Projektleitung an der H-BRS
Prof. Dr. Mike Althaus Dr. Karl Kirschner Prof. Dr. Matthias Preller Prof. Dr. Dirk Reith Prof. Dr. Jörn Oliver Sass Prof. Dr. Margit Schulze Prof. Dr. Christopher Volk Prof. Dr. Steffen Witzleben Dr. Katrin RichterDie bio-chemische Forschung ist zunehmend auf akkurate Computermodellierung und -analyse angewiesen. Dieses Forschungsfeld ist naturgemäß hoch interdisziplinär, da physikalische Grundgesetze algorithmisch umgesetzt werden müssen, um in Anwendungen der Lebenswissenschaften relevante Beiträge liefern zu können. Das Projekt und die damit verbundene Initiative UMMBAS bündelt disziplinübergreifend die starke Expertise an der H-BRS in der Methodenentwicklung, der Visualisierung und der Anwendung computergestützter Verfahren zur Entschlüsselung materialwissenschaftlicher und biochemischer Fragestellungen.
Projektleitung an der H-BRS
Prof. Dr. Matthias PrellerPublikationen
Publications since 2015 (70+ total since 1993)
- M Mobasher, AJ Hone, M Pilzecker, D Bozic, A Hecker, JM McIntosh, KN Kirschner, V Grau, RL Papke, H Andleeb, and K Richter, "Putative α7-selective ligands interact with α9-containing nicotinic acetylcholine receptors and modulate immune functions of human mononuclear phagocytes". Front. Immunol. 17 (2026) 1773637 https://doi.org/10.3389/fimmu.2026.1773637
- KN Kirschner. “Gas-Phase Benzene-Methanol Dimer Configurations: Geometries, Relative Stabilities, and Interaction Energies”. J. Phys. Chem. A 129 (2025), 8110–8119 https://doi.org/10.1021/acs.jpca.5c03923
- R Strickstrock, A Hagg, D Reith, and KN Kirschner. “Speed up Multi-Scale Force-Field Parameter Optimization by Substituting Molecular Dynamics Calculations with a Machine Learning Surrogate Model”. ChemPhysChem 26 (2025), e202500353 https://doi.org/10.1002/cphc.202500353
- R Strickstrock, A Hagg, M Hülsmann, KN Kirschner, and D Reith, “Fine-tuning property domain weighting factors and the objective function in force-field parameter optimization”. Journal of Molecular Graphics and Modelling 139 (2025), 109035 https://www.sciencedirect.com/science/article/pii/S1093326325000956
- A Hagg and KN Kirschner. “Open-Source Machine Learning in Computational Chemistry”. Journal of Chemical Information and Modeling 63 (2023), 4505–4532 https://doi.org/10.1021/acs.jcim.3c00643
- M Müller, A Hagg, R Strickstrock, M Hülsmann, A Asteroth, KN Kirschner, & D Reith. “Determining Lennard-Jones Parameters Using Multiscale Target Data through Presampling-Enhanced, Surrogate-Assisted Global Optimization”. Journal of Chemical Information and Modeling 63 (2023), 1872–1881 https://doi.org/10.1021/acs.jcim.2c01231
- W Fiedler, F Freisleben, J Wellbrock, & Kirschner, Karl N. “Mebendazole’s Conformational Space and Its Predicted Binding to Human Heat-Shock Protein 90”. Journal of Chemical Information and Modeling 62 (2022), 3604–3617 https://doi.org/10.3390/ijms221910670
- R Strickstrock, M Hülsmann, D Reith, and KN Kirschner. “Optimizing Lennard-Jones parameters by coupling single molecule and ensemble target data”. Computer Physics Communications 274 (2022), 108285 https://doi.org/10.1016/j.cpc.2022.108285
- F. Freisleben, F. Modemann, J. Muschhammer, H. Stamm, F. Brauneck, A. Krispien, C. Bokemeyer, K.N. Kirschner, J. Wellbrock, & W. Fiedler. "Mebendazole Mediates Proteasomal Degradation of GLI Transcription Factors in Acute Myeloid Leukemia," International Journal of Molecular Sciences, 2021, 22, 10670 https://doi.org/10.3390/ijms221910670
- Cesari, A.; Uccello Barretta, G.; Kirschner, K.N.; Pappalardo, M.; Basile, L.; Guccione, S.; Russotto, C.; Lauro, M. R.; Cavaliere, F. & Balzano, F. “Interaction of natural flavonoid eriocitrin with β-cyclodextrin and hydroxypropyl-β-cyclodextrin: an NMR and molecular dynamics investigation,” New J. Chem., The Royal Society of Chemistry, 2020, 44, 16431-16441 https://pubs.rsc.org/en/content/articlelanding/2020/nj/d0nj02022b#!divAbstract
- K.N. Kirschner, S. Keil, K. Seuser, and C. Siefer, “Teaching Technical Journalism with an Engineering Foundation,” in 2020 IEEE Global Engineering Education Conference (EDUCON), Porto, Portugal, 2020, pp. 808-813 https://ieeexplore.ieee.org/document/9125242 (Won Best Paper.)
- K.N. Kirschner, D. Reith, and W. Heiden, “The performance of Dunning, Jensen, and Karlsruhe basis sets on computing relative energies and geometries,” Soft Materials, 2020, 18, 200-214 https://www.tandfonline.com/doi/full/10.1080/1539445X.2020.1714656
- Schenk, M.R.; Köddermann, T.; Kirschner, K.N.; Knauer, S. & Reith, D. “Molecular Dynamics in the Energy Sector: Experiment and Modeling of the CO2/CH4 Mixture,” Journal of Chemical & Engineering Data, 2020, 65, 1117-1123 https://pubs.acs.org/doi/10.1021/acs.jced.9b00503
- Krämer, A.; Pickard, F.; Huang, J.; Venable, R.; Reith, D.; Kirschner, K.; Pastor, R. & Brooks, B. “Interactions of Water and Alkanes: Modifying Additive Force Fields to Account for Polarization Effects,” J. Chem. Theory Comput., 2019, 15 (6), 3854-3867 https://pubs.acs.org/doi/10.1021/acs.jctc.9b00016
- Mitchell, S. R.; Larkin, K.; Grieselhuber, N. R.; Lai, T.-H.; Cannon, M.; Orwick, S.; Sharma, P.; Asemelash, Y.; Zhang, P.; Goettl, V. M.; Beaver, L.; Mims, A.; Puduvalli, V. K.; Blachly, J. S.; Lehman, A.; Harrington, B.; Henderson, S.; Breitbach, J. T.; Williams, K. E.; Dong, S.; Baloglu, E.; Senapedis, W.; Kirschner, K.; Sampath, D.; Lapalombella, R. & Byrd, J. C. “Selective targeting of NAMPT by KPT-9274 in acute myeloid leukemia, Blood Advances, American Society of Hematology, 2019, 3, 242-255 https://www.bloodadvances.org/content/3/3/242
- A. Bernardi, R. Faller, D. Reith, and K.N. Kirschner, “ACPYPE update for nonuniform 1-4 scale factors: Conversion of the GLYCAM06 force field from AMBER to GROMACS,” SoftwareX, 2019, 10, 100241 https://www.sciencedirect.com/science/article/pii/S2352711018300736
- K. Kirschner, J. Bode, and D. Reith, “The International Chair - Concept and Benefits of a New Interdisciplinary Faculty Position,” in 2019 IEEE Global Engineering Education Conference (EDUCON), 2019, 775-780 https://ieeexplore.ieee.org/document/8725255
- K. N. Kirschner, W. Heiden, and D. Reith. “Small alcohols revisited: CCSD(T) relative potential energies for the minima, first- and second-order saddle points, and torsion-coupled surfaces,” ACS Omega, 3(1):419–432, 2018 https://pubs.acs.org/doi/abs/10.1021/acsomega.7b01367
- A. Bernardi, K.N. Kirschner, and R. Faller. “Structural analysis of human glycoprotein butyrylcholinesterase using atomistic molecular dynamics: The importance of glycosylation site ASN241,” PLOS ONE, 12(11):1–17, 11 2017 https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0187994
- T. Köddermann, M.R. Schenk, M. Hülsmann, A. Krämer, K.N. Kirschner, and D. Reith. “Molecular dynamics simulation of membrane free energy profiles using accurate force field for ionic liquids.” In Scientific Computing and Algorithms in Industrial Simulations. Springer, Cham, 2017 https://link.springer.com/chapter/10.1007/978-3-319-62458-7_14
- K. N. Kirschner, W. Heiden, and D. Reith. “Relative electronic and free energies of octane’s unique conformations. Molecular Physics, 115(9-12):1155–1165, 2017 https://www.tandfonline.com/doi/abs/10.1080/00268976.2016.1262076
- R. Elfgen, M. Hülsmann A. Krämer, T. Köddermann, K.N. Kirschner, and D. Reith. “Optimized atomistic force fields for aqueous solutions of magnesium and calcium chloride: Analysis, achievements and limitations,” The European Physical Journal Special Topics, 225(8):1391–1409, 2016 https://link.springer.com/article/10.1140/epjst/e2016-60112-7
- M. Hülsmann, K.N. Kirschner, A. Krämer, D.D. Heinrich, O. Krämer-Fuhrmann, and D. Reith. “Optimizing molecular models through force-field parameterization via the efficient combination of modular program packages. In Q. R. Snurr, S. C. Adjiman, and A. D. Kofke, editors, Foundations of Molecular Modeling and Simulation: Select Papers from FOMMS 2015, pages 53–77. Springer Singapore, Singapore, 2016 https://link.springer.com/chapter/10.1007/978-981-10-1128-3_4
- K.N. Kirschner, D. Reith, O. Jato, and A. Hinkenjann. “Visualizing potential energy curves and conformations on ultra high-resolution display walls,” Journal of Molecular Graphics and Modelling, 62:174–180, 2015 https://www.sciencedirect.com/science/article/pii/S1093326315300577
Weitere Infos